Bonjour à tous! Considérez les données sur les champignons, prédisez leur caractère comestible, établissez une corrélation et bien plus encore.
Nous utiliserons les données sur les champignons de Kaggle (dataframe d'origine) de https://www.kaggle.com/uciml/mushroom-classification , 2 dataframes supplémentaires seront jointes à l'article.
Toutes les opérations sont effectuées sur https://colab.research.google.com/notebooks/intro.ipynb
# e
import pandas as pd
# , confusion_matrix:
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import confusion_matrix
# :
import matplotlib.pyplot as plt
import seaborn as sns
#
mushrooms = pd.read_csv('/content/mushrooms.csv')
#
mushrooms.head()
# :
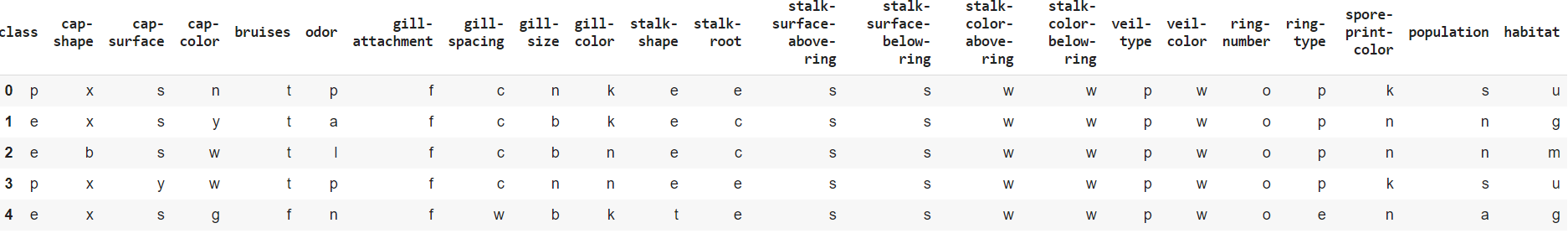
#
mushrooms.info()

#
mushrooms.shape
# LabelEncoder ( heatmap)
# ,
from sklearn.preprocessing import LabelEncoder
le=LabelEncoder()
for i in mushrooms.columns:
mushrooms[i]=le.fit_transform(mushrooms[i])
#
mushrooms.head()

# heatmap
fig = plt.figure(figsize=(18, 14))
sns.heatmap(mushrooms.corr(), annot = True, vmin=-1, vmax=1, center= 0, cmap= 'coolwarm', linewidths=3, linecolor='black')
fig.tight_layout()
plt.show()
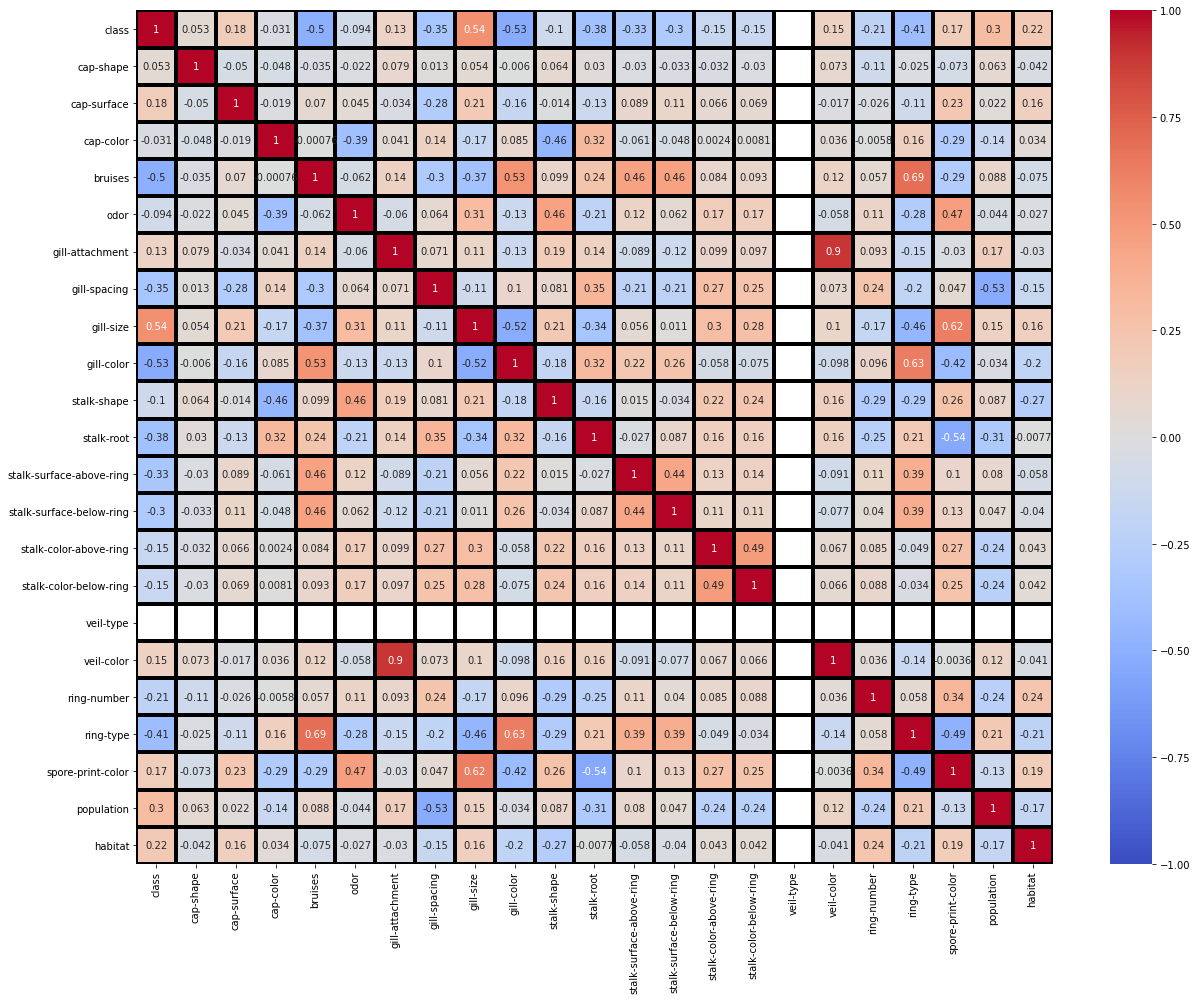
: (veil-color,gill-spacing) = +0.9 (ring-type,bruises) = +0.69 (ring-type,gill-color) = +0.63 (spore-print-color,gill-size) = +0.62 (stalk-root,spore-print-color) = -0.54 (population,gill-spacing) = -0.53 (gill-color,class) = -0.53 , . , , .
# , .
X = mushrooms.drop(['class'], axis=1)
# , .
y = mushrooms['class']
# RandomForestClassifier.
rf = RandomForestClassifier(random_state=0)
# ,
#{'n_estimators': range(10, 51, 10), 'max_depth': range(1, 13, 2),
# 'min_samples_leaf': range(1,8), 'min_samples_split': range(2,10,2)}
parameters = {'n_estimators': [10], 'max_depth': [7],
'min_samples_leaf': [1], 'min_samples_split': [2]}
# Random forest GridSearchCV.
GridSearchCV_clf = GridSearchCV(rf, parameters, cv=3, n_jobs=-1)
GridSearchCV_clf.fit(X, y)
# ,
best_clf = GridSearchCV_clf.best_params_
# .
best_clf
# confusion matrix ( ) , .
y_true = pd.read_csv ('/content/testing_y_mush.csv')
sns.heatmap(confusion_matrix(y_true, predictions), annot=True, cmap="Blues")
plt.show()

Cette matrice d'erreurs montre que nous n'avons pas d'erreurs du premier type, mais il y a des erreurs du deuxième type dans la valeur 3, ce qui pour notre modèle est un indicateur très faible tendant vers 0.
Ensuite, nous effectuerons des opérations pour déterminer le modèle avec la meilleure précision de notre df
#
from sklearn.metrics import accuracy_score
mr = accuracy_score(y_true, predictions)
#
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
#
#
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression(max_iter = 10000)
lr.fit(x_train,y_train)
#
from sklearn.metrics import confusion_matrix,classification_report
y_pred = lr.predict(x_test)
cm = confusion_matrix(y_test,y_pred)
#
log_reg = accuracy_score(y_test,y_pred)
#K
#
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors = 5, metric = 'minkowski',p = 2)
knn.fit(x_train,y_train)
#
from sklearn.metrics import confusion_matrix,classification_report
y_pred = knn.predict(x_test)
cm = confusion_matrix(y_test,y_pred)
#
from sklearn.metrics import accuracy_score
knn_1 = accuracy_score(y_test,y_pred)
#
#
from sklearn.tree import DecisionTreeClassifier
dt = DecisionTreeClassifier(criterion = 'entropy')
dt.fit(x_train,y_train)
#
from sklearn.metrics import confusion_matrix,classification_report
y_pred = dt.predict(x_test)
cm = confusion_matrix(y_test,y_pred)
#
from sklearn.metrics import accuracy_score
dt_1 = accuracy_score(y_test,y_pred)
#
#
from sklearn.naive_bayes import GaussianNB
nb = GaussianNB()
nb.fit(x_train,y_train)
#
from sklearn.metrics import confusion_matrix,classification_report
y_pred = nb.predict(x_test)
cm = confusion_matrix(y_test,y_pred)
#
from sklearn.metrics import accuracy_score
nb_1 = accuracy_score(y_test,y_pred)
#
plt.figure(figsize= (16,12))
ac = [log_reg,knn_1,nb_1,dt_1,mr]
name = [' ',' ',' ',' ', ' ']
sns.barplot(x = ac,y = name,palette='colorblind')
plt.title(" ", fontsize=20, fontweight="bold")

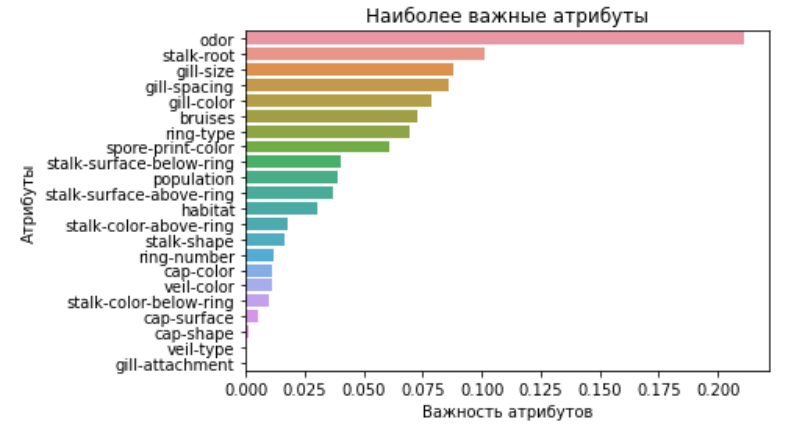
Nous pouvons conclure que le modèle le plus précis pour nos prédictions est un arbre de décision.